Enhancer-Promoter RNA Interaction Atlas
What's EP-Atlas?
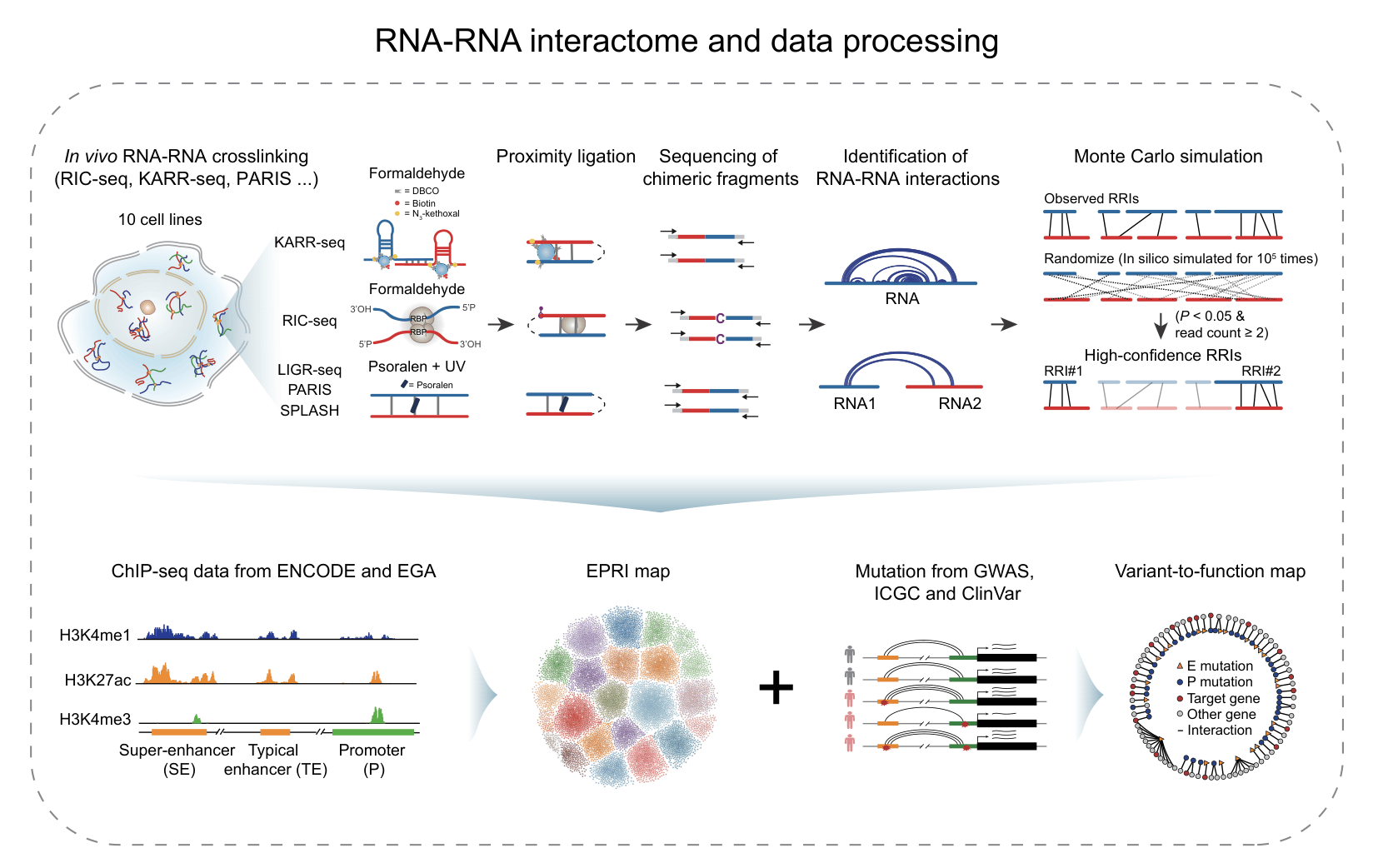
Enhancers and promoters are fundamental cis-regulatory elements that orchestrate precise spatiotemporal control of gene expression. Increasing evidence has demonstrated that enhancer–promoter communication is mediated not only through DNA-based chromatin looping, but also through RNA-associated interactions, highlighting an additional regulatory layer beyond classical three-dimensional genome organization. Recent advances in high-throughput RNA-RNA interactome technologies, such as RIC-seq and KARR-seq, have enabled direct, transcriptome-wide detection of RNA-mediated chromatin interactions. These approaches provide unique insights into the dynamic and context-dependent nature of enhancer–promoter regulation. However, the resulting datasets remain scattered across studies, processed using heterogeneous pipelines, and are often difficult to integrate and systematically explore.
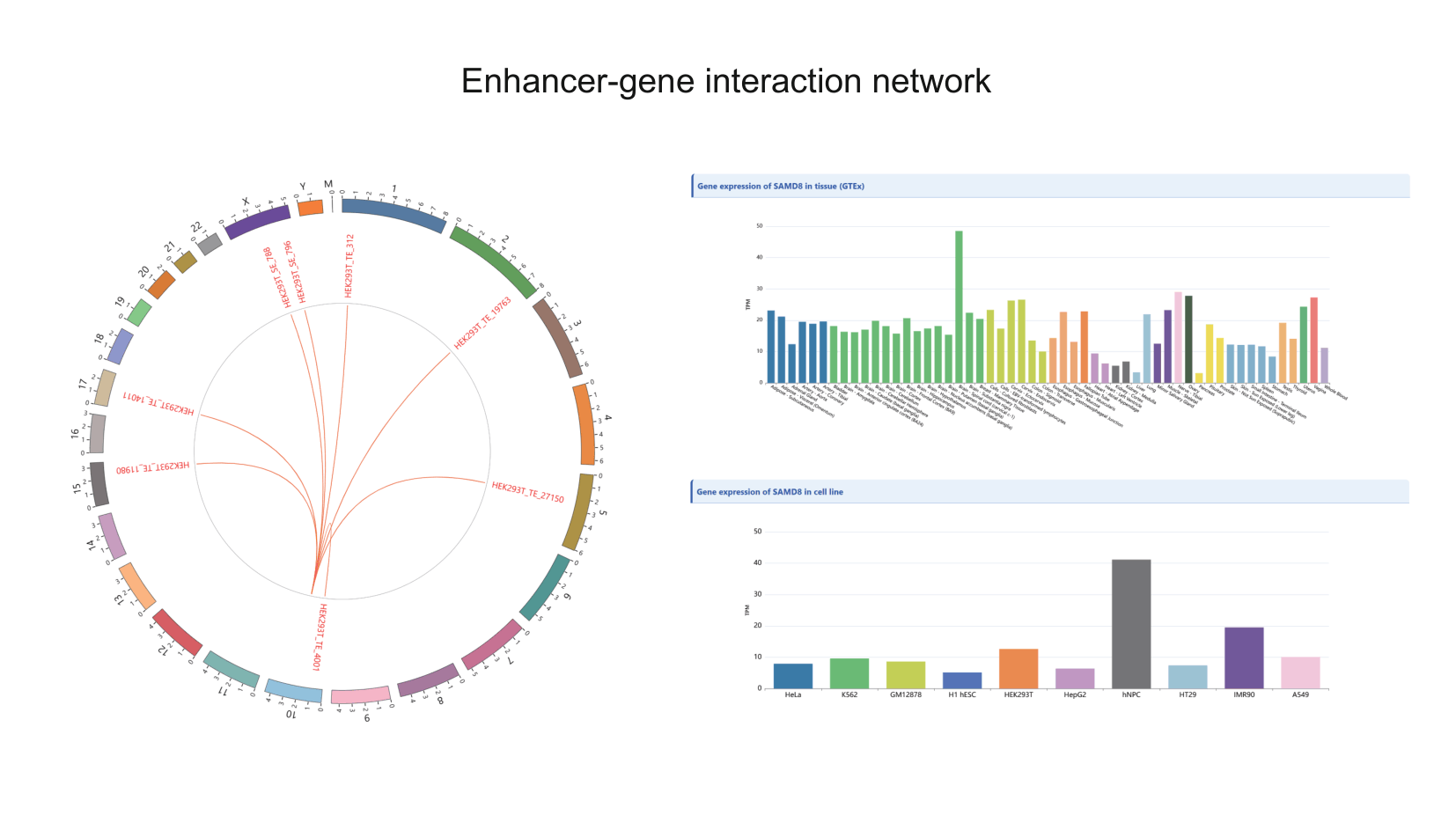
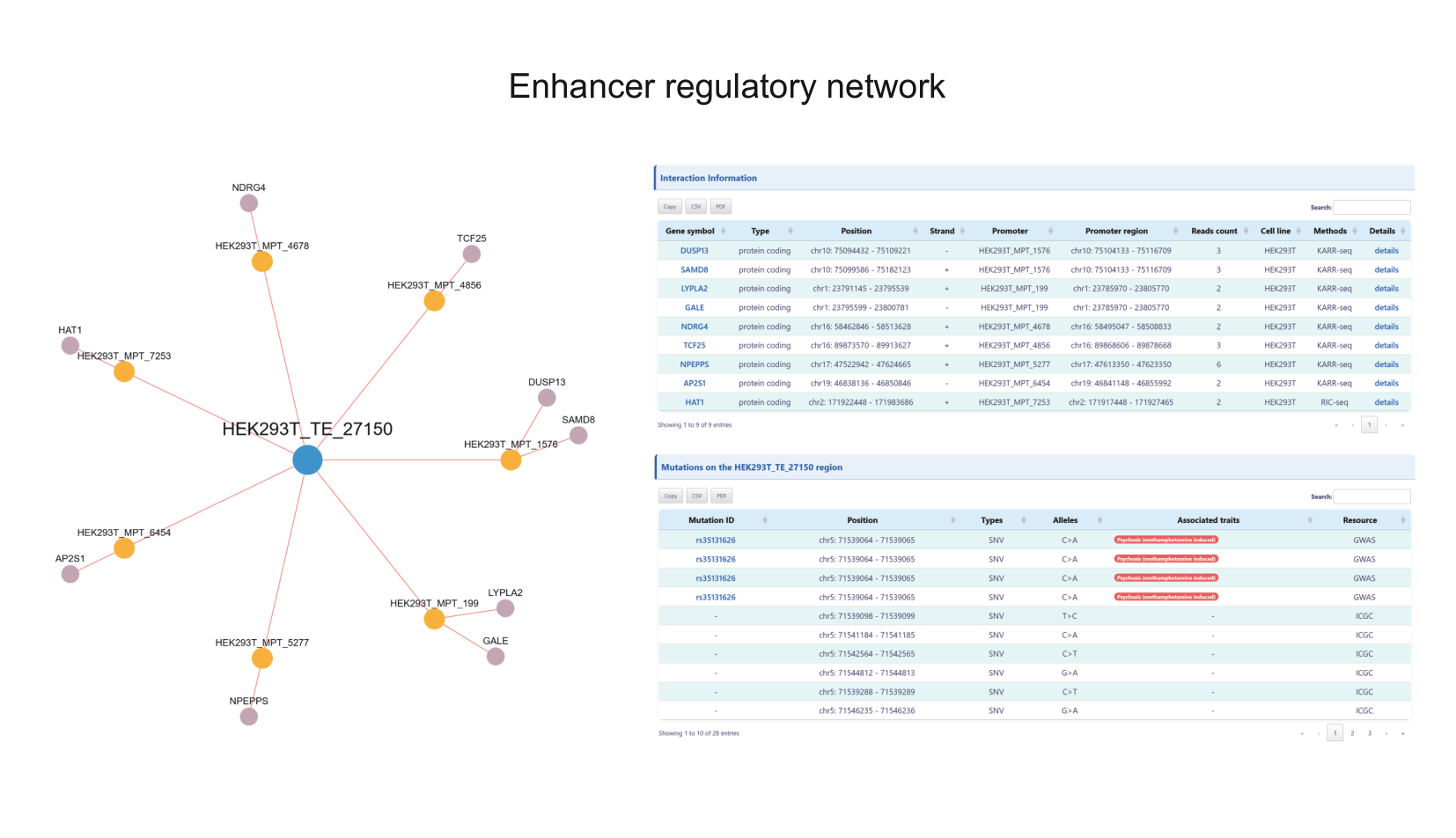
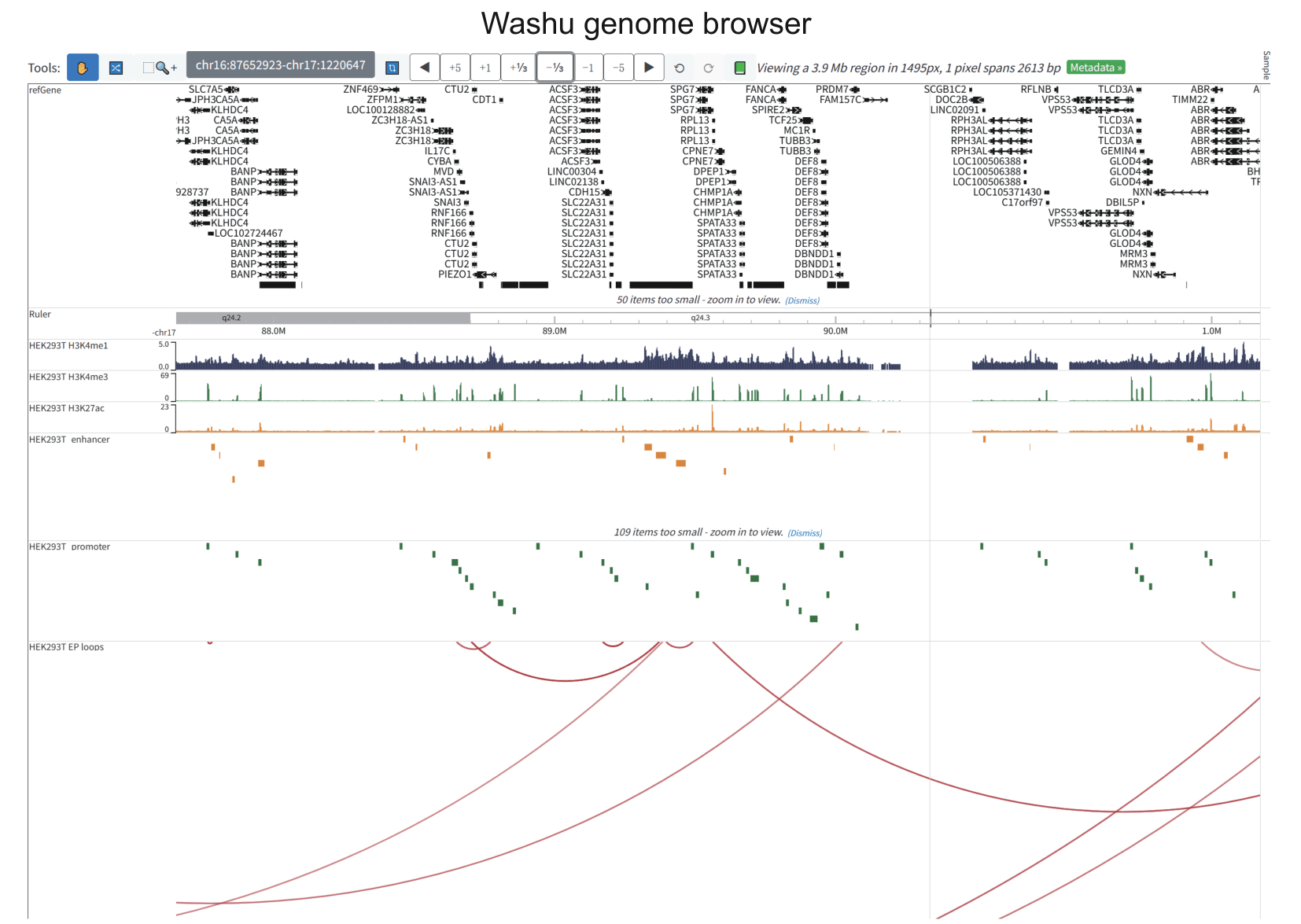
To address these limitations, we developed EP-Atlas (https://epatlas.cn), a comprehensive database that integrates high-throughput datasets from RIC-seq, KARR-seq, LIGR-seq, SPLASH, and PARIS to systematically identify 980,639 high-confidence enhancer–promoter RNA interactions (EPRIs). Each interaction is extensively annotated with gene information, expression profiles, genomic interaction regions, supporting chimeric read counts, chromatin states, as well as genetic variants from GWAS Catalog, ClinVar, and ICGC, together with expression quantitative trait loci (eQTLs) from PanQTL v2 and ncRNA-eQTL databases. EP-Atlas provides a unique platform for investigating cell-type-specific, RNA-mediated enhancer–promoter regulatory networks and for interpreting the functional impact of noncoding variation in human traits and disease.
Contact us
Principal Investigator: Yuanchao Xue
State Key Laboratory of Epigenetic Regulation and Intervention, Institute of Biophysics, Chinese Academy of Sciences, Beijing 100101, China
Email: ycxue@ibp.ac.cn